LECTURE 1
A Broad Perspective
- Structural biology
o Drug discovery
o Small molecule design
- Comparative genomics
o Sequence analysis
- Evolutionary biology
- Gene regulation – OUR FOCUS
= big data analysis, statistics and mathematical modelling, machine learning
Structural Bioinformatics
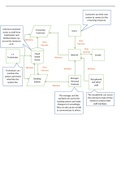
- Design of a small molecule to redirect the trajectory of drug resistance
o Trimethoprim (TMP) derivative impedes antibiotic resistance evolution
§ Targets DHFR (bacterial enzyme required for DNA synthesis in a
bacteria)
• Repeated treatment of the bacteria with TMP to drive
mutations leading to resistance
o Found that L28R mutation (leucine to arginine) à
structural change in DHFR à decreased aSinity of
TMP for DFHR
§ Most common mutation that can drive
resistance
§ Can model with molecular dynamics
§ Altered chemistry of TMP à 4’DTMP (binds
well to mutated DFHR)
• TMP and 4’DTMP both work on WT
bacteria
• Mutant bacteria (L28R) à TMP less
eSective than 4’DTMP at killing
bacteria
§ Drives selection of alternative mutations that confer drug
resistance but also decrease catalytic capacity of DFHR (reduces
fitness)
• Eg c-35t mutation
Comparative Genomics
- Identification of functional elements in what appear to be random sequences
- Design of an RNA therapeutic to target this element to treat rare genetic epilepsy
- Evolutionary conservation is associated with function
o Human and mouse HSP90 à highly conserved aa sequence
o Highly conserved sequences lead to highly conserved proteins with
conserved structures upon folding
, § Therefore, drug design needs to target the slight diSerences in
order to target one organism’s protein but not another (eg target
parasite HSP90 and not human)
- But… long non coding RNA à eg CHASERR IncRNA = low levels of conservation
§ Have syntenic counterparts but low sequence conservation
o CHASERR =
§ Transcribed upstream of Chd2
• Has positional conservation throughout vertebrates
§ Regulates CHD2 expression in cis
• Essential for mouse viability
• Knockout of CHASERR à increased CHD2 production
o Negative regulator
§ Chd2 haploinsuSiciency in humans à epilepsy and intellectual
disability
§ Chromatin remodeler
• Regulates H3.3 deposition
• Essential to neurogenesis and myogenesis
§ How is the function of CHASERR encoded in these rapidly evolving
sequences?
• Some lncRNAs will have multiple short conserved elements
dispersed throughout sequence
o Goal is to find a combination of short elements
§ Order of conserved motifs is important
• Why look for conserved combinations
of short elements and not just any
short element?
o Probability of the single
sequence occurring by chance
is high while probability of
specific sequences in a
specific order has a lower
probability of occurring by
chance à possibly more
functional
- If 1 = allowed
- If 0 = not allowed
- Want to maximize sum of non
intersecting lines
- Found conserved sequence in
last exon of CHASERR
, Gene Regulation
- Identification of genes that drive regulation of genes of interest
o RNA-seq experiments!
§ Quantify expression levels of all genes post neuronal injury – what
genes are upregulated
LECTURE 2
Back to Basics
- DNA = double helix that comprises two anti parallel strands
- Polymerase can only transcribe in 5’ to 3’ direction
o Can transcribe from wither of the two parallel strands
- Assembly of genome has + and – strand
o Important for alignment etc
- Always read 5’ to 3’ and the coding direction is always 5’ to 3’
RNA Seq
- Sequencing the transcriptome
- Massively parallel RNA sequencing
- EG can compare between two conditions
o Before and after injury
§ Compare the transcriptome in both conditions
• Which genes are upregulated
- How is a gene expressed at the RNA level
o Extract RNA from cells
o Reverse transcribe RNA to fragments of dsDNA
§ Why fragments?
• DiSicult to get RT enzyme to run along the whole length of
RNA
o IneSicient transcription
o Sequence dsDNA molecules using illumine sequencing technology
§ Using flow cell and fluorescence
- Overview of RNA seq
o Isolation of total RNA
o Enrichment of non ribosomal RNA
§ Ensures that the majority of reass to not correspond to a small
number of rRNAs rather than mRNAs or lncRNAs (95-98% of all
human RNAs may be rRNAs)
• polyA selection à enrich for mature (spliced)
polyadenylated RNAs
• rRNA depletion
o Conversion of RNA to cDNA
o Construction of fragment library
o Sequencing platform
o Generation of single or paired end reads
o Alignment and assembly of reads
o Downstream analysis