Yeast Genetics
Characteristics
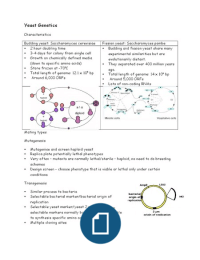
Budding yeast: Saccharomyces cerevisiae Fission yeast: Saccharomyces pombe
2 hour doubling time Budding and fission yeast share many
3-4 days for colony from single cell experimental similarities but are
Growth on chemically defined media evolutionarily distant.
(down to specific amino acids) They separated over 400 million years
Store frozen at -70oC ago.
Total length of genome: 12.1 x 106 bp Total length of genome: 14 x 106 bp
Around 6,000 ORFs Around 5,000 ORFs
Lots of non-coding RNAs
Mating types
Mutagenesis
Mutagenise and screen haploid yeast
Replica plate potentially lethal phenotypes
Very often – mutants are normally lethal/sterile – haploid, no need to do breeding
schemes
Design screen – choose phenotype that is viable or lethal only under certain
conditions
Transgenesis
Similar process to bacteria
Selectable bacterial marker/bacterial origin of
replication
Selectable yeast marker/yeast 2 micron origin
selectable markers normally based on mutants unable
to synthesis specific amino acid
Multiple cloning sites
, Cloning by functional complementation
ARG2 gene – gene involved in arginine
biosynthesis pathway
Infect yeast cells with a library of genes
(read notes)
Targeted Mutagenesis
Homologous recombination extremely efficient in yeast
Only 25bp of homologous sequence required
Targeting constructs and systematic gene disruption
- Recombination between homologous regions 1 and 2 replaces
MFG1 coding sequence with URA3 cassette
- Constructs generated very efficiently for any gene by PCR
Genome-wide tagged mutation libraries
All 6000 yeast ORFs have been targeted and tagged
4500 ORFS not required for viability under standard
conditions – screen for phenotypes for competitive
ability in yeast –loci that confer increased fitness
- Place 10 cells of each of the 6000 strains in a flask –
subject flask to various stressful conditions
- Tagged strains can be subjected to competition and
% of each strain determined by DNA extraction and
microchip analysis
Ageing of pool maintained in stationary phase containing 3000 S. pombe mutants
Maintain yeast cells at stationary phase – don’t grow but can die
Samples collected at various time points – DNA extraction, PCR, sequencing of
barcodes follows
Sequencing data; no. of reads per tag/mutant – determine presence and proportion
of mutants in different samples
Epistasis Analysis
Characteristics
Budding yeast: Saccharomyces cerevisiae Fission yeast: Saccharomyces pombe
2 hour doubling time Budding and fission yeast share many
3-4 days for colony from single cell experimental similarities but are
Growth on chemically defined media evolutionarily distant.
(down to specific amino acids) They separated over 400 million years
Store frozen at -70oC ago.
Total length of genome: 12.1 x 106 bp Total length of genome: 14 x 106 bp
Around 6,000 ORFs Around 5,000 ORFs
Lots of non-coding RNAs
Mating types
Mutagenesis
Mutagenise and screen haploid yeast
Replica plate potentially lethal phenotypes
Very often – mutants are normally lethal/sterile – haploid, no need to do breeding
schemes
Design screen – choose phenotype that is viable or lethal only under certain
conditions
Transgenesis
Similar process to bacteria
Selectable bacterial marker/bacterial origin of
replication
Selectable yeast marker/yeast 2 micron origin
selectable markers normally based on mutants unable
to synthesis specific amino acid
Multiple cloning sites
, Cloning by functional complementation
ARG2 gene – gene involved in arginine
biosynthesis pathway
Infect yeast cells with a library of genes
(read notes)
Targeted Mutagenesis
Homologous recombination extremely efficient in yeast
Only 25bp of homologous sequence required
Targeting constructs and systematic gene disruption
- Recombination between homologous regions 1 and 2 replaces
MFG1 coding sequence with URA3 cassette
- Constructs generated very efficiently for any gene by PCR
Genome-wide tagged mutation libraries
All 6000 yeast ORFs have been targeted and tagged
4500 ORFS not required for viability under standard
conditions – screen for phenotypes for competitive
ability in yeast –loci that confer increased fitness
- Place 10 cells of each of the 6000 strains in a flask –
subject flask to various stressful conditions
- Tagged strains can be subjected to competition and
% of each strain determined by DNA extraction and
microchip analysis
Ageing of pool maintained in stationary phase containing 3000 S. pombe mutants
Maintain yeast cells at stationary phase – don’t grow but can die
Samples collected at various time points – DNA extraction, PCR, sequencing of
barcodes follows
Sequencing data; no. of reads per tag/mutant – determine presence and proportion
of mutants in different samples
Epistasis Analysis